Why from v0.2 to v0.69? Cause I’m immature AF and it probably made you want to read this blog post.
Back in October 2018, I released a tool called BeaconGraph after attending the SANS SEC617: Wireless Penetration Testing and Ethical Hacking course taught by James Vidal. I released a PoC of BeaconGraph after I realized that airgraph-ng could use a more modern look. However, that version of BeaconGraph was not very user-friendly, as it was more of a Proof-of-Concept than a usuable tool. Additionally, the UI was pretty trash because I lack adequate web design skills, and attempted to write all of the HTML, CSS and Javascript myself with Cytoscape as the rendering engine. It looked like this:
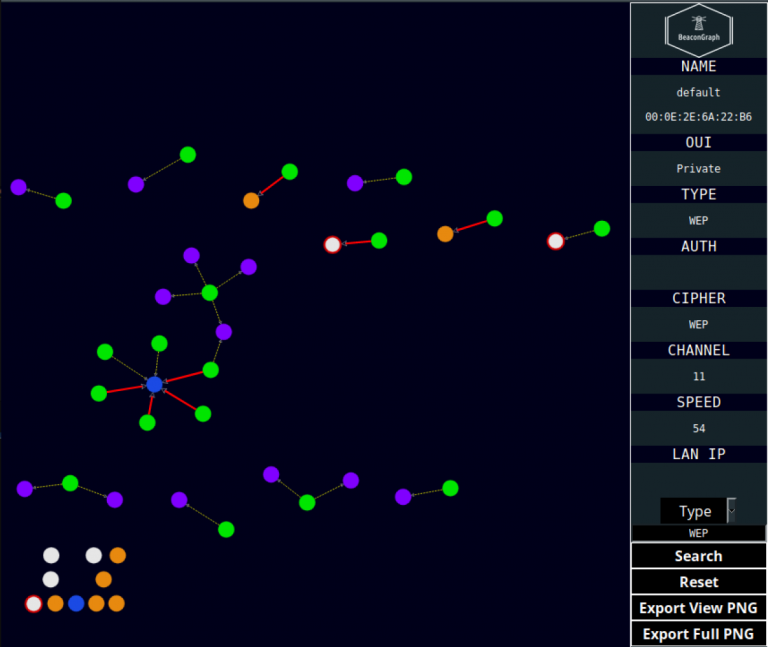
While it was a great attempt to write my own code, I realized at the time that adding additional functionality or even just changing around the UI was going to be a pain. When I finally decided to buckle down and work on it again, I discovered that the team over at plot.ly had written an open source dashboard framework called Dash and had recently implemented a version of Cytoscape. This was perfect, as I now had everything I need to basically rewrite BeaconGraph from scratch. With this, I am able to announce the release of BeaconGraph v0.69!
You can find in on my Github here: BeaconGraph
What’s New
PyQt5
Look at that login screen. Just look at it.
Besides moving over to a Dash framework, the other major change to BeaconGraph is that it is now displayed using PyQt5. This eliminates the need to open a browser, although since it is a Flask app, you can open a browser to the page and navigate to the page manually if you want. This creates a more GUI-friendly app and in the future will be compiled into binaries for releases on different OSes. Additionally, this will allow for a greater extension of UI control beyond Dash when it is necessary.
Dash UI
By far the most important part of the update, Dash provides a clean looking interface in Python that avoids having to deal with a lot of the underlying hassle for HTML/CSS/JS. I found that creating this UI was not only simpler than trying to write everything myself, but the built-in components and the use of callbacks was everything that was needed to take BeaconGraph to the next level.
Additionally, since all the command line arguments have been stripped (for now), you can upload Airodump CSVs using the upload button in the upper left corner. This UI is a work in progress, so for now, all output of any functionality can only be seen in the command line. In the future, there will be a way to view command line output within the app.
The database tab in the upper right corner will display all of the database stats. However, the legend on the bottom left will display the numbers that are currently loaded on screen. In a future update, you will be able to set the node colors to whatever you want, which is great for colorblind users.
The nodes tab displays information about the currently selected node. Additionally, it also includes a search feature that allows to query the database by any of the properties shown. You can even select multiple of the same property to show multiple nodes!
Currently, the Queries tabs and Settings tabs are empty. The Settings tab will contain settings such as database URL, color changes, etc. The Queries tab will be filled with queries that aren’t as easy to display via the search bar. In a way in the future update, you will have the ability to write your own queries to display data.
Parsing changes
I didn’t have this type of data before when I originally starting writing but now BeaconGraph properly displays mesh networks. This is great for showing wireless APs that are connected to each other. Previously, the nodes were separated and it wasn’t immediately clear that the separate APs were part of the same mesh. Here’s an example of a Sonos sound system mesh network.
In this display, we can see that WPA APs are associated with each other, indicating that it is a mesh network.
What’s Gone
Node focus
Unfortunately, since Dash Cytoscape does not yet implement it, the ability to focus in on a particular node in the display has been lost. For now, this functionality can somewhat be expressed by the search functionality and manual zoom.
Node highlight
Although the highlight feature was a great addition, it was not part of the core Cytoscape functionality. Hopefully in the future, this can be re-implemented, particularly when a bunch of relationships are displayed on screen.
Known Bugs
Random display movement
There is a weird case where sometimes the nodes will keep jumping around the screen and move further and further
away from each other. Currently, the solution is to simply restart the app. However, unless this is not a Dash
Cytoscape bug I’m not aware of, it’s likely due to the high parameters nodeOverlap and nodeRepulsion
features that are set for the Cytoscape display. I am not sure if it is currently possible, but I will look for
a way to make this number more reasonable and dynamic based on the amount of nodes currently displayed in view.
Future updates
There are a couple of things I’d like to add before the official v1 release, such as settings and queries. Beyond that however, I would like to implement additional input sources such as probemon or Kismet. It was suggested to me that I should implement some sort of compatibility with Bettercap. This is also likely to happen in the future. :)
Acknowledgements
Thanks to the following people for providing input and feedback on design and neo4j implementation: